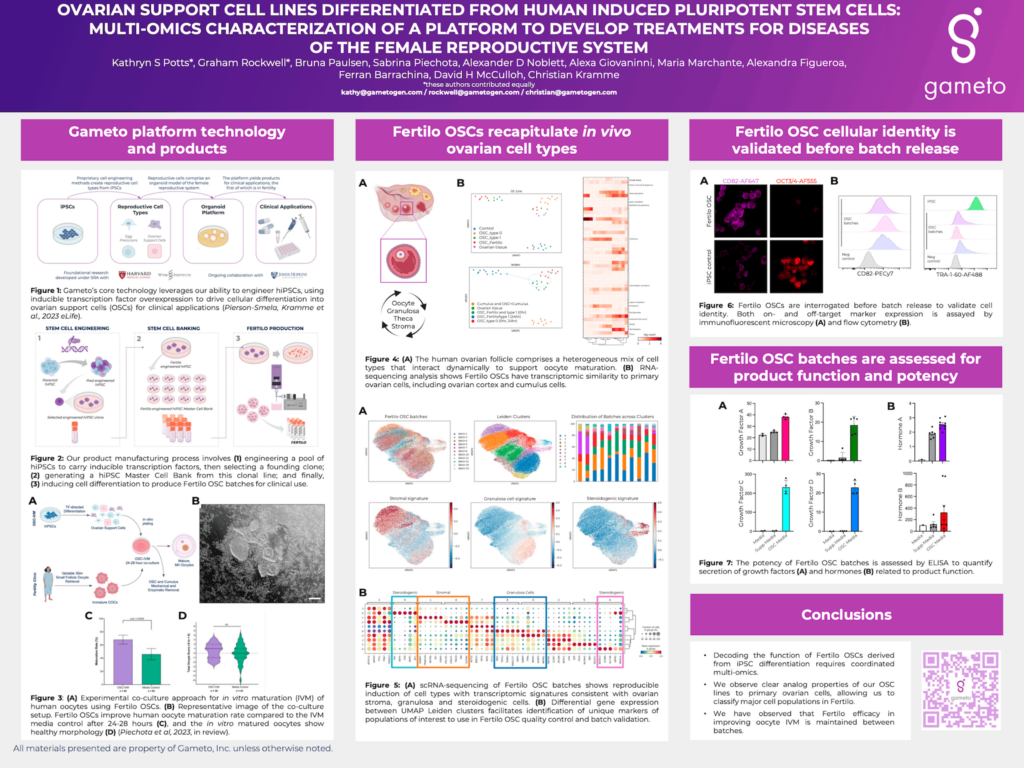
An in vitro model of the human ovary would advance our understanding of female sexual development and reproduction, enabling the development of novel fertility treatments such as robust oocyte in vitro maturation technology. A crucial functional component of such a model are ovarian support cells (OSCs),including granulosa, theca and stromal cell lineages. Together, the diverse milieu of OSCs are responsible for shaping ovarian signaling and the in vivo oocyte maturation environment.
We recently reported an efficient five day protocol to generate heterogenous OSCs from human induced pluripotent stem cells (hiPSCs) through transcription factor (TF) overexpression. We have demonstrated their ability to recapitulate dynamic ovarian function in vitro including follicle formation and steroidogenesis to support in vitro oocyte maturation. Here we characterize the diversity of our OSCs through a broad multi-omics analysis approach of multiple batches of differentiated cells. Using high throughput single cell RNAseq we identify distinct and overlapping transcriptomic profile populations of cells amongst different induction batches. Comparing profiles to signature genes of known ovarian cell populations from the literature and primary ovarian tissue transcriptomics, we identified sub-populations with strong similarity to stromal, pre-granulosa and granulosa lineages.
We also bring together metabolomic, proteomic and ELISA analyses of our cell line batches to investigateOSC function including paracrine signaling. Our findings are elucidating the OSC mechanism of action and feed into improved quality assurance and control metrics for OSC batch production. Overall we are able to construct a robust discovery process and quality control for the production of consistent function profiles in hiPSC-derived OSCs.